 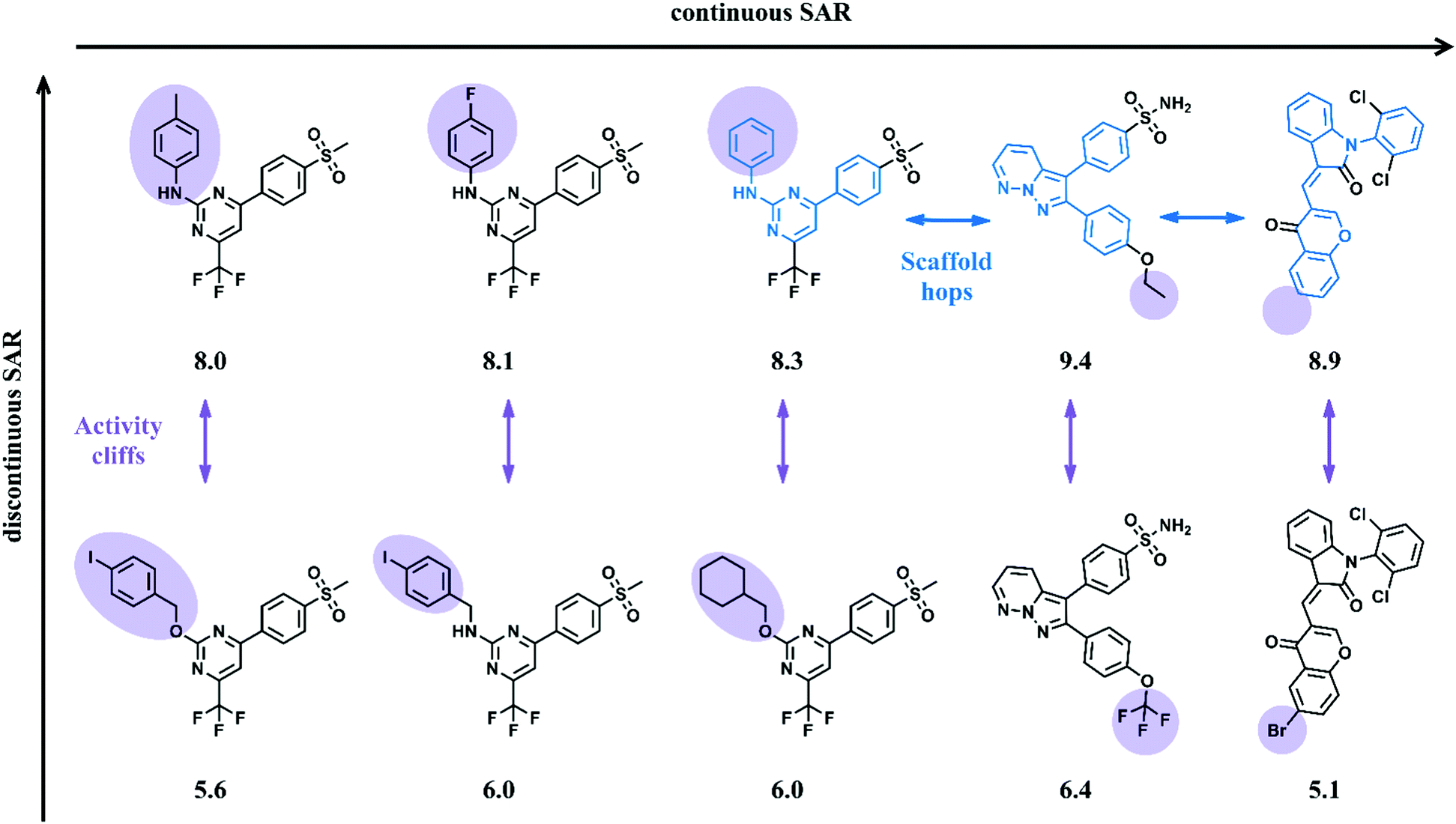
Some 3D approaches are more explicit than others - docking versus CoMFA for example. 3D approaches on the other hand are generally more explicitly informative, in the sense that one can directly understand the nature of ligand - receptor interactions that underlie an observed SAR. For example, statistical QSAR approaches bsed on 2D descriptors that ignore stereochemistry can be miss key elements of an SAR that depend on chirality. It is important to realize that the choice of modeling technique can influence the to what extent and in how much detail an SAR can be explored. For a comprehensive review of QSAR methodologies the reader is referred to previous reviews of the field. We can broadly divide them into two groups - those based on statistical or data mining methods (e.g., regression models) and those based on physical approaches (e.g., pharmacophore models). Over the last 60 years there have been a multitude of ways to capture structure-activity relationships. Section 5 discusses some approaches to exploring SAR data that do not involve explicit model development and finally Section 6 provides a summary of this topic. 
Section 4 discusses the role of SAR databases. Section 3 discusses recent work in the area of structure-activity landscapes and how they can be used as an alternative view of SAR data. Section 2 provides an overview of common modeling techniques that are used to encode SAR relationships. The remainder of the text is structured as follows. In addition to considering modeling algorithms, I will also briefly discuss the use of databases as a source of SAR data and how they can be used to inform and enhance the exploration of SAR trends. 
The goal of this chapter is to highlight the different type of SAR modeling methods, and specifically, how they support the task of exploring chemical space to elucidate and optimize structure-activity relationships in a drug discovery setting. These methods cover the spectrum in terms of complexity and utility. That is, a reduced or simplified representation of reality, replete with assumptions and limitations. Fundamentally, SAR models are just that: models. While it is certainly true that computational methods can help in identifying, explaining and predicting structure-activity relationships (SAR), it is also true that naïve usage (or even misuse) of these techniques can lead to misleading results. Rather, they can provide a guide to the experienced user by integrating and summarizing large amounts of pre-existing data to suggest useful structural modifications. It should be kept in mind that computational methods do not replace medicinal chemistry domain knowledge. Coupled with in silico modifications of structures, one can easily prioritize large screening decks or even generate new compounds de novo and ascertain whether they belong to the SAR being studied or not. These methods allow one to build a variety of models to capture and encode one or more SARs, which can then be used to predict activities for new molecules. How does one rapidly identify the most promising series amongst them? In these scenarios, in silico methods allow rapid and efficient characterization of SARs. For example, in a primary HTS it is possible that one is faced with hundred of chemical series. While the intuition and experience of a medicinal chemist is vital to these efforts, the data generated by modern high throughput experimental techniques can overwhelm the capabilities of a single chemist. For example, most lead optimization projects will try and improve potency, reduce toxicity and ensure sufficient bioavailability, amongst other properties. Invariably, the development of a chemical series involves optimizing multiple physicochemical and biological properties simultaneously. Fundamentally, an understanding of the SAR for a set of molecules, allows one to rationally explore chemical space, which, in the absence of “sign posts” is essentially infinite. Working with SAR starts from identifying whether an SAR actually exists in a collection of molecules and their associated activities to trying to elucidate the details of one or more SARs and subsequently using that information to make structural modifications to optimize some property or activity. Structure-activity relationships (SAR) are key to many aspects of drug discovery, ranging from primary screening to lead optimization. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
